Cytometry team
Cell Secrets Unveiled
The number of cells, markers and samples measured per cytometry experiment keeps increasing, making traditional analysis methods, such as manual gating on 2D scatter plots, infeasible in many cases. The Cytometry Team of the SaeysLab develops algorithms and pipelines for an automated analysis of this high-throughput single-cell technique.
We focus on the one hand on algorithm development, with tools like PeacoQC for quality control, CytoNorm for batch effect removal and FlowSOM for clustering.
On the other hand, we also have several ongoing collaborations with clinicians to build complex but explainable analysis pipelines, studying diseases including primary immunodeficiencies (PID), acute myeloid leukemia (AML) and non-small cell lung cancer (NSCLC). Our goal here is to bring these pipelines to the clinic.
Highlighted cytometry papers
Analyzing high-dimensional cytometry data using FlowSOM
Katrien Quintelier, Artuur Couckuyt, Annelies Emmaneel, Joachim Aerts, Yvan Saeys & Sofie Van Gassen
Nature Protocols 2021
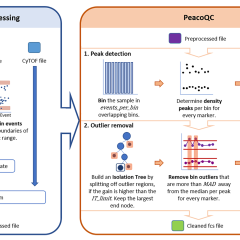
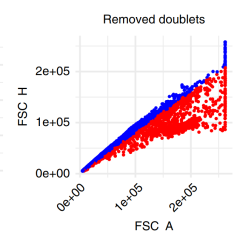
PeacoQC: Peak-based selection of high quality cytometry data
Annelies Emmaneel, Katrien Quintelier, Dorine Sichien, Paulina Rybakowska, Concepción Marañón, Marta E. Alarcón-Riquelme, Gert Van Isterdael, Sofie Van Gassen & Yvan Saeys
Cytometry Part A. 2021
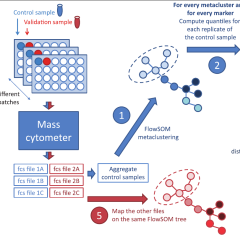
CytoNorm: A Normalization Algorithm for Cytometry Data
Sofie Van Gassen, Brice Gaudilliere, Martin S. Angst, Yvan Saeys & Nima Aghaeepour
Cytometry Part A. 2019
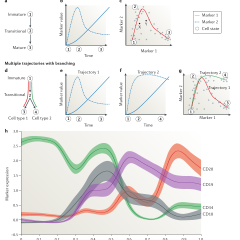
Computational flow cytometry: helping to make sense of high-dimensional immunology data
Yvan Saeys, Sofie Van Gassen & Bart N. Lambrecht
Nature reviews immunology 2016
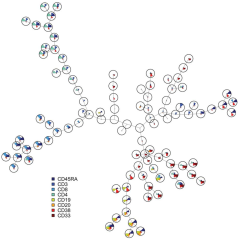
FlowSOM: Using self-organizing maps for visualization and interpretation of cytometry data
Sofie Van Gassen, Britt Callebaut, Mary J Van Helden, Bart N Lambrecht, Piet Demeester, Tom Dhaene & Yvan Saeys
Cytometry Part A. 2015

